AluHunter is a suite of computer programs that automate the task of finding potentially polymorphic Alu elements for use in primate phylogenetics.
Below is more information on the project.
- Alu Elements in Systematics
- The Old Way of Finding Alu Elements
- Bioinformatic Identification of Potentially Polymorphic Alu Elements
- How AluHunter Works
- How You Can Use These Data
- Technical Details
Alu Elements in Systematics
Alu elements are a class of primate-specific SINEs. Throughout evolutionary time they've spread throughout the genome in a copy-and- paste mechanism. These insertions have two characteristics that make them ideal for use in primate systematics: they are nearly free of homoplasy and are of a known ancestral state. This means you can use Alu presence to distinguish between different primate evolutionary lineages. If two taxa share an Alu and a third does not, you know almost certainly that those first two taxa are more closely related to each other than they are to the third.
 The
great thing about Alu elements is that once they've been
identified, further fine-scaled screening just requires PCR and
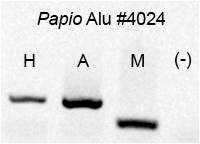
gel electrophoresis. To the right is a photograph of one such gel. The
PCR amplification primers were designed from sequence flanking an
Alu insertion. So if an Alu is present in a sample, the amplicon is
roughly 300 basepairs longer than if the Alu is absent. You can see
in the gel that this particular Alu insertion is present in
Hamadryas (H) and Anubis (A) baboons, but absent in the rhesus
macaque (M).
The
great thing about Alu elements is that once they've been
identified, further fine-scaled screening just requires PCR and
gel electrophoresis. To the right is a photograph of one such gel. The
PCR amplification primers were designed from sequence flanking an
Alu insertion. So if an Alu is present in a sample, the amplicon is
roughly 300 basepairs longer than if the Alu is absent. You can see
in the gel that this particular Alu insertion is present in
Hamadryas (H) and Anubis (A) baboons, but absent in the rhesus
macaque (M).
Below is the sequence from a macaque (top) aligned with that of a Hamadryas baboon (bottom). Note the Alu-sized gap and the conserved flanking regions where a primer can sit.

Since the Alu is present in two species of baboon, but not in the macaque, we can infer that this Alu was inserted on the lineage leading to baboons since the split with macaques, possibly in one of the following scenarios:
|
|
|
|
Given a large enough number of Alu elements, each grouping some genera, species, or populations together, you can see how you could infer an evolutionary tree for all primates. Because of their low levels of homoplasy, this Alu-based tree might more closely represent the true evolutionary relationships than phylogenies based on DNA sequence similarity or shared morphological characters. Alu elements could resolve phylogenies that have proven difficult or impossible to systematists thus far, and an Alu that groups a population of interest could be a useful marker in population genetics.
The Old Way of Finding Alu Elements
Unfortunately, useful Alu elements are difficult to find. Alu elements that are fixed in your taxon of interest are phylogenetically useless. Previous methods of identifying potentially polymorphic Alu elements involved cloning Alu sequences, designing primers, and then screening the Alu elements in your taxon of interest. Since the ages of many Alu insertions are older than your taxon of interest, many Alu elements screened this way turned out to be fixed. It was a costly and time-consuming process.
Bioinformatic Identification of
Potentially Polymorphic Alu Elements
Say you're using lab-based methods to find Alu elements that are variable in baboons. You'll find many that are fixed in primates, fixed in Old World monkeys and apes, or fixed in the African Papionins. But if you can screen out the ones that are also present in a sister taxon, you'd be left with only those Alu elements that were inserted in the lineage of interest after the two groups shared a least common ancestor. So if you take all Alu elements found in baboons and ignore those that are also present in macaques, you'll be left with only those that were inserted on the baboon lineage since the two groups split. These are the potentially polymorphic Alu elements that may be useful in resolving phylogeny in your taxon of interest, in this case baboons.

Given an Alu found in baboon sequence, you need to determine if it is present in the orthologous outgroup sequence. The easiest way to do so is to take the flanking regions around the baboon Alu and BLAST them against the macaque genome. If both flanks have matches in the macaque genome with a roughly 300 basepair long gap in between (see image above, right), the Alu is inferred to be present in baboons and macaques, and therefore inserted before the lineages split. If the flanks are present in the macaque, but with no gap in between (image above, left), the Alu is inferred to be present in baboons, but not in macaques and was thus inserted since the two lineages split. This Alu is potentially phylogenetically informative.
Below is the outcome of BLASTing the baboon Alu in the gel photo above against the macaque genome. The right and left flanks, which were separated by an Alu insertion in the original Hamadryas baboon sequence, are found right next to one another in the macaque.
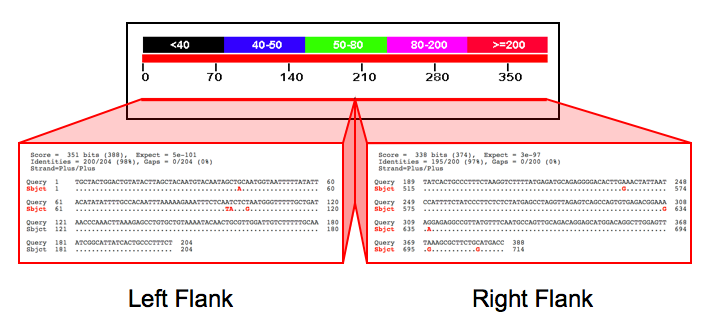
These potentially phylogenetically informative Alu elements can then be screened via PCR in the taxa that have diverged since the split with the outgroup. So an Alu found in a baboon sequence but absent in the macaque can then be screened in the African Papionins. The Alu above was originally found in the Hamadryas baboon and further screening via PCR showed it groups all the African Papionins.
How AluHunter Works
AluHunter is a suite of computer programs that automate the task of finding potentially polymorphic Alu elements for use in primate phylogenetics. Broadly, the algorithm performs the following steps:
- Once every 24 hours, AluHunter downloads all new DNA that's been sequenced for a given taxon and submitted to NCBI GenBank.
- It searches the DNA for Alu insertions and adds all Alu elements to a database.
- It then grabs the flanking regions that surround the Alu insertion in the source DNA.
- Next AluHunter BLASTs the flanks against an outgroup genome or genomes, and decides if an Alu is present, absent, or indeterminate.
- For those Alu elements that are absent in the outgroup genome, it designs PCR primers from the flanks surrounding the Alu.
- All interesting Alu element's data and primers are uploaded into an online database, accessible through this website.
AluHunter searches GenBank nightly for new primate sequences that it hasn't screened yet, scans those sequences for Alu elements, and screens them against an appropriate outgroup. The actual method is fairly simple; the novelty of AluHunter lies in its degree of automation and its scale. The end result is a real-time database of all Alu elements that have been sequenced and submitted to GenBank.
How You Can Use These Data
This website has been designed to facilitate use of the data that AluHunter generates. To use the Alu primers, you only need a lab capable of PCR and gel electrophoresis, and samples in your taxon of interest. (We've successfully typed Alu elements in DNA from low quality sources.) If you meet those requirements, simply click on "Find Alus by Taxon" above and select your taxon of interest. AluHunter will pick the nearest primate genome to use as an outgroup and generate a list of Alu elements that may be polymorphic within your taxon. It can also give you PCR amplification primers for testing for the presence of those Alu elements. The primers can be exported as a spreadsheet or text document for easy ordering with any of the major custom oligonucleotide companies.
If you are interested in using the Alu primers generated by AluHunter in your research, please cite the latest publication. If you have any questions, send me a message via the contact page above.
There is a lot of sequence in which to search for Alu elements and new primate genomes to screen them against. More Alu elements are being found nightly. If you would like to be alerted when Alu elements for your taxon of interest are found, please send me a message via the contact form.
Technical Details
AluHunter is written mostly in Perl, and uses BioPerl, RepeatMasker, BLAST, and Primer3. A shell script written in bash controls the automated execution of the code. Screening the Alu elements against outgroup genomes is the most computationally intensive part of the program and takes places either locally or on NYU's high performance computing cluster, depending on the size of the job. Automated primer design occurs for interesting Alu elements, and a MySQL database is built nightly and automatically populated with the potentially phylogenetically informative Alu elements.











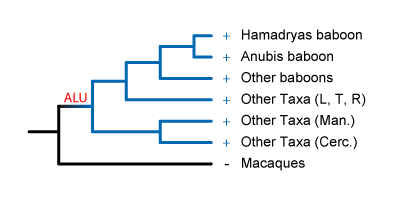 The Alu could have been
inserted right after the baboon lineage split from the macaque lineage. It's then
fixed in all African Papionins.
The Alu could have been
inserted right after the baboon lineage split from the macaque lineage. It's then
fixed in all African Papionins.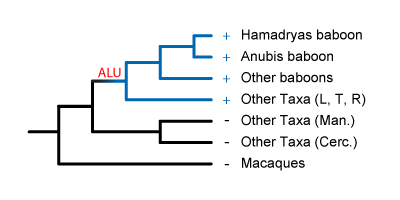 Inserted a
bit later, it would be present in some African Papionins, but not others.
These Alus would be useful for systematics within the African Papionins, like for resolving
mangabey phylogeny.
Inserted a
bit later, it would be present in some African Papionins, but not others.
These Alus would be useful for systematics within the African Papionins, like for resolving
mangabey phylogeny.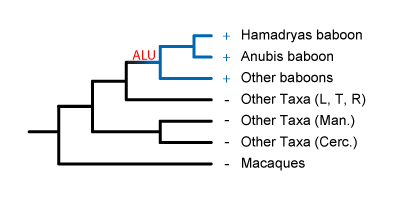 It could have been inserted
later still, and present just in the baboon genus Papio. These Alus would
be useful for identifying members of this genus.
It could have been inserted
later still, and present just in the baboon genus Papio. These Alus would
be useful for identifying members of this genus.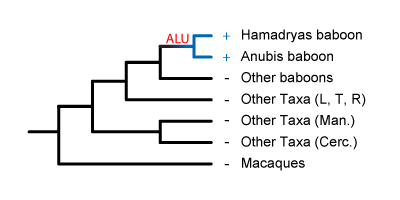 The most recent insertions would
be present in some members of Papio, but not others. If the Alu is polymorphic within
the genus, it can tell us something about species-level phylogeny of the baboons.
The most recent insertions would
be present in some members of Papio, but not others. If the Alu is polymorphic within
the genus, it can tell us something about species-level phylogeny of the baboons.